marie julie fave
ph.d student, montreal
e-mail: marie-julie.fave@mail.mcgill.ca
Time has passed since the great evolutionary synthesis of the XXth century and modern biology has made formidable advances in many fields, from paleontology to genomics. There is an ongoing movement to revise the evolutionary theory framework in the light of these new findings and integrating more recent research fields such as EvoDevo or epigenetics. Far from claiming that I will solve the issue myself, my research focuses on integrating the central field of modern evolutionary theory, population genetics, with the more recently emerged field of EvoDevo.
Specifically, I am interested in the molecular and developmental mechanisms underlying the parallel evolution of similar traits in independent populations. Recent work on traditional or emerging model organisms have shown that closely related species can evolve independently similar phenotypes in response to similar ecological pressures, but simultaneously exhibit divergent molecular mechanisms underlying the trait formation. Characterizing such variation and trying to decipher the relative contribution of natural selection, genetic drift or gene flow to create and maintain this variation in developmental processes is the main focus of my research.
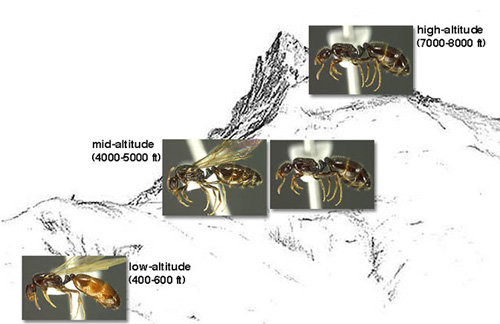
I am conducting my work on a system of populations of the ant Monomorium emersoni that inhabits the sky island archipelago in Southeastern Arizona, mountains separated from each other by a widespread desert since the last glaciation. Interestingly, queens of this species have evolved a wing polymorphism where wingless queens are to be found in higher proportions at higher elevations. This pattern is repeated on the different genetically isolated mountain ranges, and we propose that divergence has evolved in the wing patterning gene network in wingless queens belonging to different isolated mountain populations.
Queen castes of the Monomorium species-complex along an altitudinal gradient:
Publications:
Suen et al. (2011) The Genome Sequence of the Leaf-Cutter Ant Atta cephalotes Reveals Insights into Its Obligate Symbiotic Lifestyle, PLoS Genetics, In Press.
Smith et al. (2011) The Draft Genome of the Globally Widespread and Invasive Argentine ant (Linepithema humile), Proceedings of the National Academy of Sciences, In Press.
Smith et al. (2011) A draft genome of the red harvester ant, Pogonomyrmex barbatus: a model for reproductive division of labor and social complexity, Proceedings of the National Academy of Sciences, In Press.
Favé and Col (In prep) eFECTIV: Shape analysis using elliptical harmonics. Available under Programs, Protocols and Datasets.
Favé and Turgeon (2008) Patterns of genetic diversity in Great Lakes Bloaters (Coregonus hoyi) with a view to future reintroduction in Lake Ontario, Conservation Genetics (9), 281-293
Favé, Duchesne and Turgeon (2008) Inbreeding dynamics in reintroduced, aged-structured populations of highly fecund species, Consevation Genetics (9), 39-48
Turgeon and Favé (2006) Identification of a genetically diverse and compatible source of bloater (Coregonus hoyi) for reintroduction in Lake Ontario, Completion report for the Great Lakes Fisheries Commission.
